User Manual
This manual describes how to access data and use the tools on the Genome Database for Rosaceae (GDR). Please use the sidebar on the left to navigate to different parts of the manual, or click on the section titles below. You can access the next page of the manual by clicking on the title of the next page below.
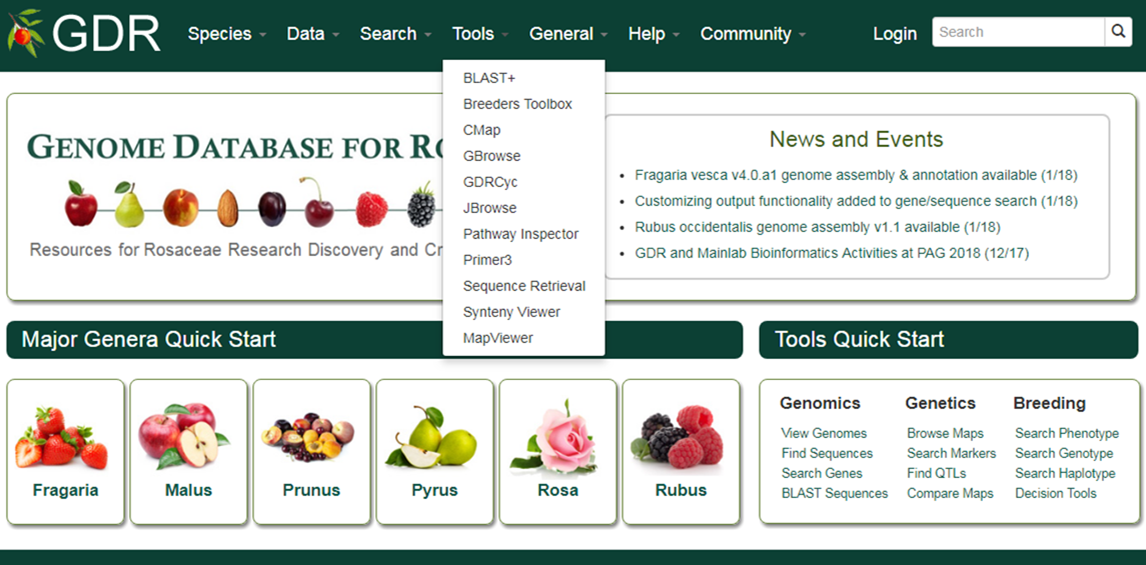
Homepage Overview
The GDR home page can be reached at www.rosaceae.org
For quick access to data, visit the Species, Data or Search section in the navigation bar or click appropriate link from the Major Genera Quick Start or Tools Quick Start section.
.png)
Species Overview Page
A species page is available for main species, genera and entire Rosaceae family so that users can easily access the data and tools for the species of interest. Species pages can be accessed under the 'species' pull-down menu in the main navigation bar.
.png)
The species pages have a resources bar on the left panel so that users can quickly access data and tools for the species. Depending on the species, more or less items will be displayed on the resource bar. Below are some items in the resource bars.
1. Genome page
Where whole genome sequences are available, hyperlinks to each genome assembly page are shown on the resources bar.
2. GDR Cyc Pathways
Where PlantCyc databases are available for a species, hyperlinks to each PlantCyc page are shown on the resources bar. The predicted genes from the whole genome sequences were utilized in the construction of PlantCyc (metabolic pathway) databases using PathwayTools. Currently (September 2013), three PlantCyc databases, PeachCyc, AppleCyc and FragariaCyc, are available in GDR.
3. Unigene page
Each species page has a hyperlink to the corresponding unigene page for the genus. Unigenes are constructed for the entire Rosaceae family and for each genus (Prunus, Malus, Fragaria, Rosa, and Pyrus) using the publicly available Rosaceae ESTs downloaded from dbEST at NCBI.
4. Maps
Each species page has hyperlink to dynamically generated list of the genetic maps that are available in GDR.
5. Whole Genome
A whole genome page provides summary information on the available whole genome data for the species.
6. Links
The links subpage shows various useful links outside GDR for the species.
7. KEGG Analysis Reports
The predicted genes from whole genome sequences and EST unigenes are associated with KEGG pathway terms. The link in the resources bar leads to the KEGG analysis reports page where users can view a Venn diagram with GO terms and the number of associated sequences, browse the GO term heirarchy, choose a term and view/download all the associated sequences.
8. Germplasm
The germplasm page provides a list of germplasm that are stored in GDR.
9. GO Analysis Reports
The predicted genes from whole genome sequences and EST unigenes are associated with Gene Ontology terms. The link in the resources bar leads to the GO analysis reports page where users can view, seach and download the GO associated data.
If you have any questions/comments/feedback about the species site, please let us know via the contact form.
Data Searches
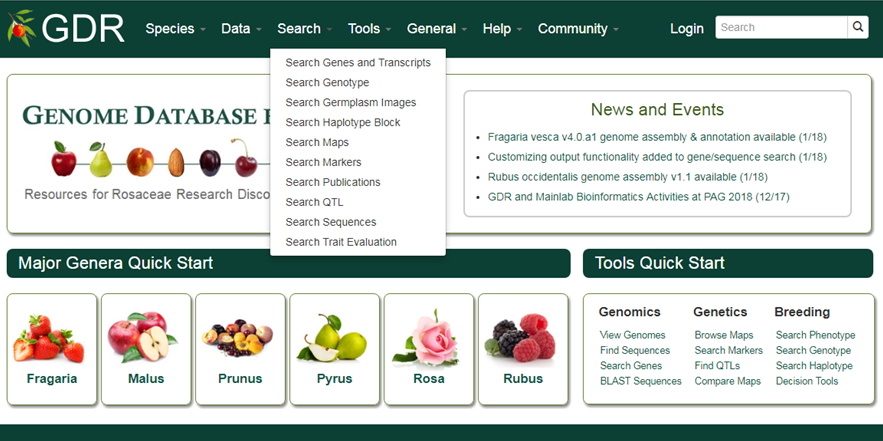
Search Genes and Transcripts
Search Genes and Transcripts is a page where users can search for genes and transcripts from various datasets available in GDR. Users can search for genes from various datasets: predicted genes from whole genome assemblies,a single non-redundant list of Rosaceae genes with gene symbols (GDR Gene Database), or gene and mRNA sequences parsed out from NCBI nucleotide database. These genes and mRNAs parsed out from NCBI sequences are aligned to the reference whole genome sequences when possible. When expert-contributed information is available, these gene names are associated directly with the predicted genes from whole genome assembly. Users can also search from transcripts from RefTrans sets, reference transcriptome sets built from all publicly available transcripts, or EST unigene contigs. For more detail, refer to 'Description of Gene and Transcript Dataset' page.
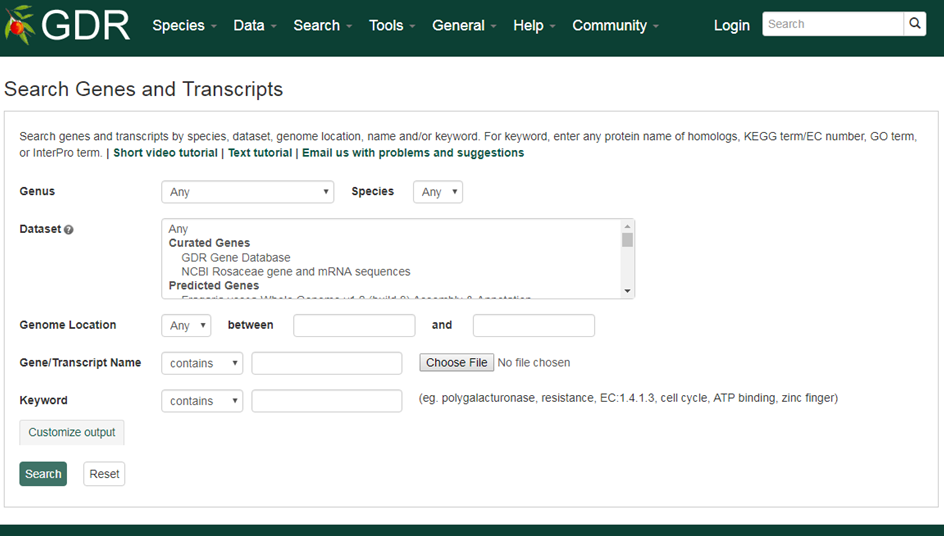
Please Note: All the search categories below, except the file upload, can be combined.
1. Genus/Species
Use these drop-down menus to limit results to genes from a specific genus or species. Once a genus is chosen, species for the genus are dynamically populated in the species dropdown. Major genera are listed on top of the Genus list under 'Common Selections' and the rest are listed as 'All options' in alphabetical order.
2. Dataset
Use this drop-down menu to limt the results to sequences from a specific dataset. Go to 'Description of Sequence Dataset in GDR' for more details. Multiple options can be selected by holding down the "Ctrl" key.
3. Genome Location
Users can limit their results of predicted genes by their genome location. When a genome assembly is chosen in the drop-down menu next to 'Dataset', the corresponding chromosome or scaffold names are dynamically displayed in the 'Genome Location' drop-down menu. Choose any option and then type in the position in bp in the text boxes.
4. Gene/Transcript Name
Users can search genes and transcripts by name for an exact match, contains, starts with or ends with the input, by selecting the desired option from the drop-down menu. The search is case-insensitive. Example gene names are MADS1, LFY2, or ppa027130m. Gene or transcript names in a file, separated by a new line, can be uploaded to do a batch search.
3. Keyword
Users can limit their result by associated funcitonal terms. Predicted genes from whole genome assembly and transcripts have been annotated with some of the followings: homology to genes of closely related or plant model species, InterPro protein domains, GO terms, KEGG pathway and ortholog terms. Users can enter any protein name (eg. polygalacturonase), KEGG term/EC number (eg. resistance, EC:1.4.1.3), GO term (eg. cell cycle, ATP binding), or InterPro term (eg. zinc finger) in the text box to limit the results with the entries that are associated with the functional annotation terms.
If you have any questions/comments/feedback about this search page, please let us know via the contact form.
Search Genotype
Search SSR Genotype is a page where users can search the SSR genotype data by dataset name, marker name, germplasm name and/or species. Click the next tab to search for SNP Genotype. To search for SSR genotype data only for cultivars and breeding selections please visit the 'Search Genotyping Data' page in the Breeders Toolbox.
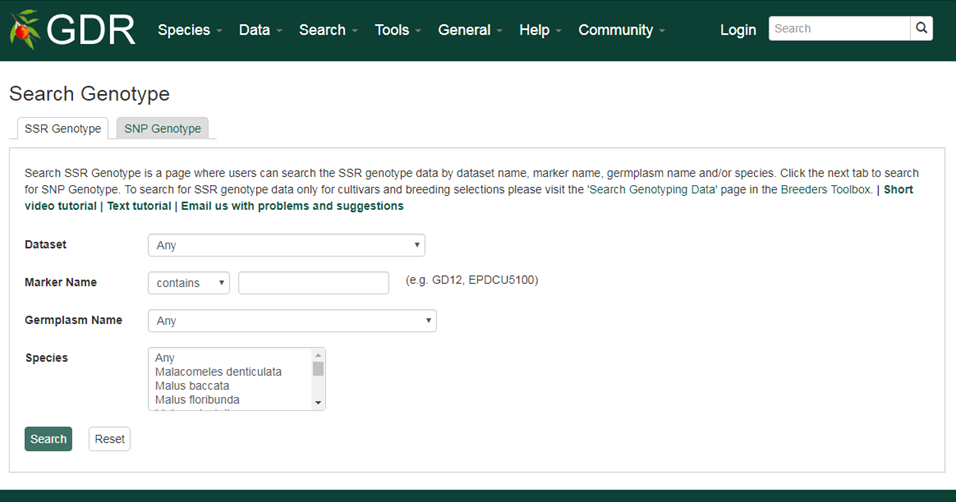
Please Note: All the search categories below can be combined.
1. Dataset
Users can search SSR genotype data by dataset from a dropdown list.
2. Marker Name
Users can search for SSR genotype data that used a specific marker by typing the marker name in the text box. Users can search for an exact match, contains, starts with or ends with the input, by selecting the desired option from the drop-down menu.
3. Germplasm Name
Users can search for SSR genotype data from a specific germplasm by choosing from the drop-down menu.
4. Species
Users can search for SSR genotype data from a specific species by choosing from the drop-down menu.
Search SNP Genotype is a page where users can search for the SNP genotyope dataset based on the germplasm and SNP markers used in the dataset. Click the next tab to search for SSR Genotype. To search for SNP genotype data only for cultivars and breeding selections please visit the 'Search Genotyping Data' page in the Breeders Toolbox.

Please Note: All the search categories below can be combined.
1. Dataset
Users can search SNP genotype data by dataset from a dropdown list.
2. Species
Users can search for SNP genotype dataset from a specific species by choosing from the drop-down menu.
3. Germplasm Name
Users can search for SNP genotype dataset from a specific germplasm by choosing from the drop-down menu.
4. SNP
Users can search for SNP genotype dataset that used a specific marker by typing the marker name in the text box. Users can search for an exact match, contains, starts with or ends with the input, by selecting the desired option from the drop-down menu.
5. Genome
Users can search SNP genotype data by the anchored position of SNPs in one of the whole genome sequences. Choose a genome in the drop-down menu next to 'Genome' then the corrposponding chromosome or scaffold names will by dynamically generated in the 'Chr/Scaffold' drop-down menu. Choose any option and then type in the position in bp in the text boxes.
If you have any questions/comments/feedback about this page, please let us know via the contact form.
Search Germplasm Images
Search Germplasm Images is a page where users can search for germplasm images available in GDR. The search can be restricted by genus, species, germplasm name and the legend of the image.
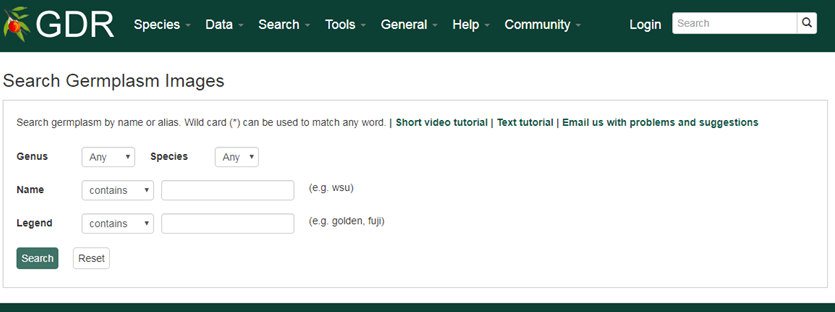
Please Note: All the search categories below can be combined.
1. Genus/Species
Use these drop-down menus to limit results to genes from a specific genus or species. Once a genus is chosen, species for the genus are dynamically populated in the species dropdown. Major genera are listed on top of the Genus list under 'Common Selections' and the rest are listed as 'All options' in alphabetical order.
2. Name
Users can search images by germplasm name for an exact match, contains, starts with or ends with the input, by selecting the desired option from the drop-down menu. The search is case-insensitive.
3. Legend
Users can limit their result by associated legend of the image. Most images of Malus species have pedigree in their legends so users can search images by entering parent's name in the legend.
If you have any questions/comments/feedback about this search page, please let us know via the contact form.
Search Haplotype Block
Search Haplotype Block is a page where you can search for haplotype blocks, a genomic region which was identified to have a distinct combination of SNP genotype. The sesarch options include species, halotype block name or genome location to which the haplotype block is aligned. From the individual haplotype block page, you can view all the haplotype (alleles) identified along with the SNP genotypes that constitutes each genotype.
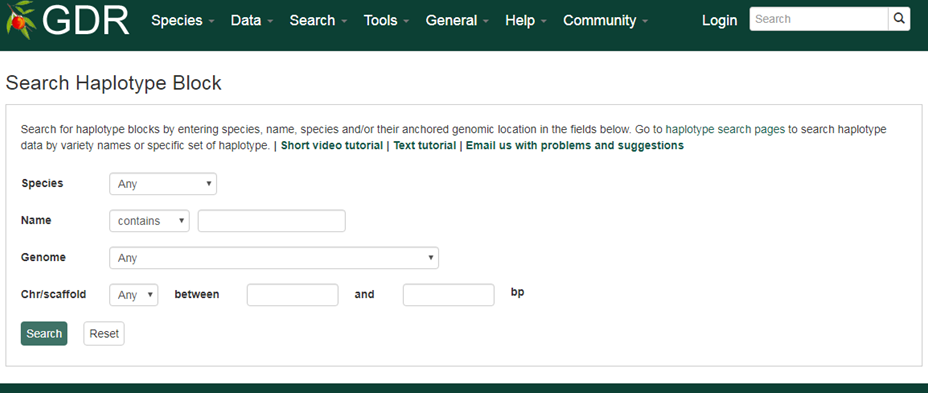
Please Note: All the search categories below can be combined.
1. Species
Use these drop-down menus to limit results to a specific species.
2. Name
Users can search by haplotype block name for an exact match, contains, starts with or ends with the input, by selecting the desired option from the drop-down menu. The search is case-insensitive.
3. Genome
Users can limit their results of haplotype blocks by their aligned genome location. When a genome assembly is chosen in the drop-down menu next to 'Genome', the corresponding chromosome or scaffold names are dynamically displayed in the 'Chr/scaffold' drop-down menu. Choose any option and then type in the position in bp in the text boxes.
If you have any questions/comments/feedback about this search page, please let us know via the contact form.
Search Maps
Search Maps is a page where users can find and view Maps by species.
.png)
1. Species
Users can search for Maps from a specific species by choosing from the drop-down menu.
If you have any questions/comments/feedback about the marker search sites, please let us know via the contact form.
Search Publications
Search Publications is a page where users can search for publication using a combination of keywords (in the abstract or title), all or partial titles, authors, and other categories. Search results link to the publication detail pages that contain the abstract, citation, external link to the full article, and other details. The GDR houses information about publications on Rosaceae genomics, genetics, and breeding research. Details about publications were imported to the GDR from NCBI PubMed and the USDA National Agricultural Library using the query: (abstract: trait OR QTL OR gene OR genome OR map OR microsatellite OR annotation OR EST OR marker OR seqeuence) AND (abstract: rosaceae or prunus or pyrus or fragaria or malus or rubus). Additionally, details of publications from other journals not present in these databases are added.
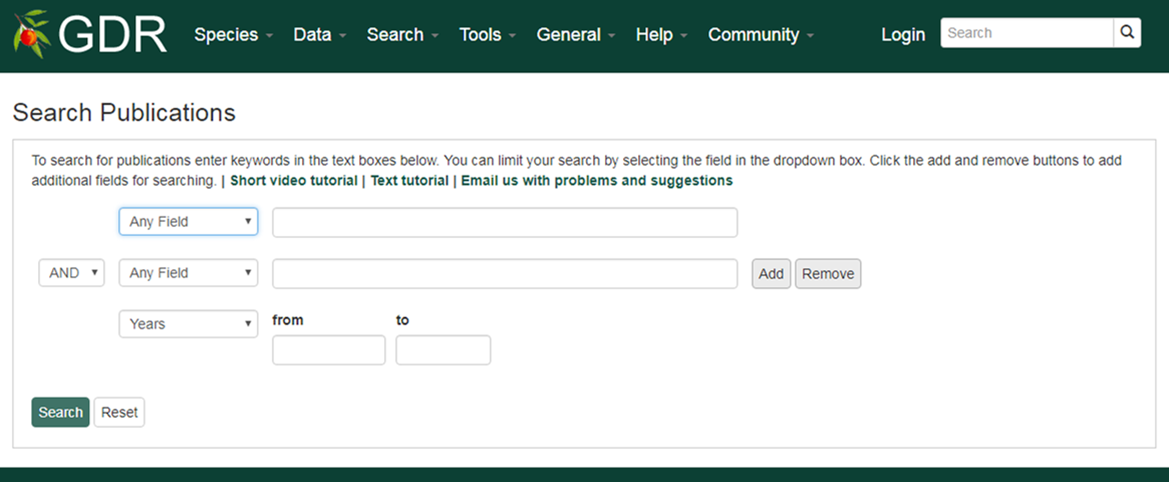
1. Users can select a field in the drop-down menu (abstract, authors, citation, journal name, title or year) and the type in keywords in the textbox
2. Users can expand their query by choosing 'AND, OR, NOT' and add additional field.
3. Users can click the plus symbol to add additional field.
4. Users can type in years in xxxx format limit the results between certain years.
If you have any questions/comments/feedback about the publications site, please let us know via the contact form.
Search QTL
Search QTL is a page where users can search for Quantatitive or Qualitative (Mendelian) Trait Loci in GDR.
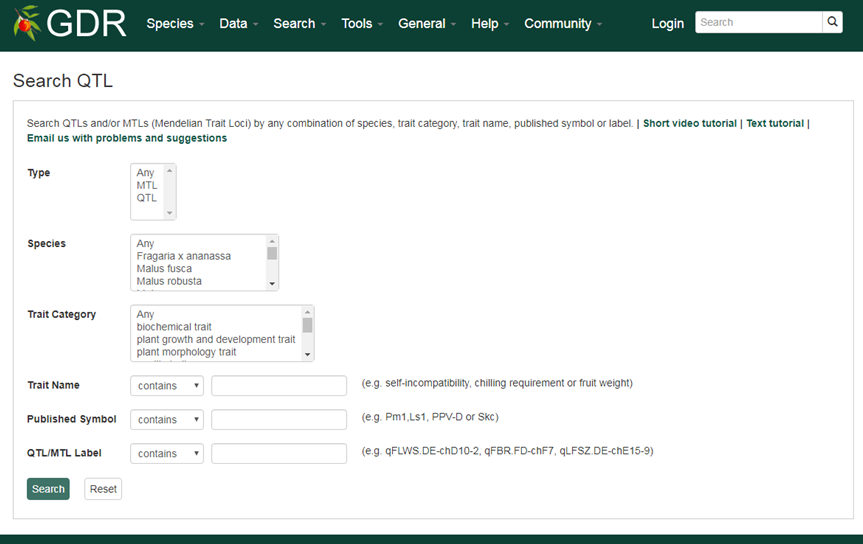
Please Note: All the search categories below can be combined.
1. Type
Users can search trait loci by type, QTL or MTL.
2. Species
Users can search trait loci by species by choosing one of the options displayed in the drop-down menu. Users can choose multiple options by holding down the "Ctrl" key.
3. Trait Category
Trait loci are associated with one or more terms of eight trait categories (anatomy and morphology, biochemical, growth and development, quality, stature or vigor, sterility or fetility, stress and yield). Users can choose multiple options by holding down the "Ctrl" key.
4. Trait Name
Trait loci are associated with trait names that belongs to the Rosaceae Trait Ontology. The Rosaceae Trait Ontology contains extra terms that are necessary for the description of Rosaceae traits in addition to the terms in Trait Ontology (developed with a focus on the traits of grass species). Examples include self-incompatibility, chilling requirement or fruit weight. Users can search these fields for an exact match, contains, starts with or ends with the input, by selecting the desired option from the drop-down menu. Users can also search by aliases. The search is case-insensitive.
5. Published Symbol
Published symbols can be used to search for trait loci. Examples include Pm1,Ls1, PPV-D or Skc. Users can search these fields for an exact match, contains, starts with or ends with the input, by selecting the desired option from the drop-down menu.
6. QTL/MTL Label
Search by the QTL/MTL label given by the GDR team.
An example QTL qSI.TE-ch5.1 indicates that:
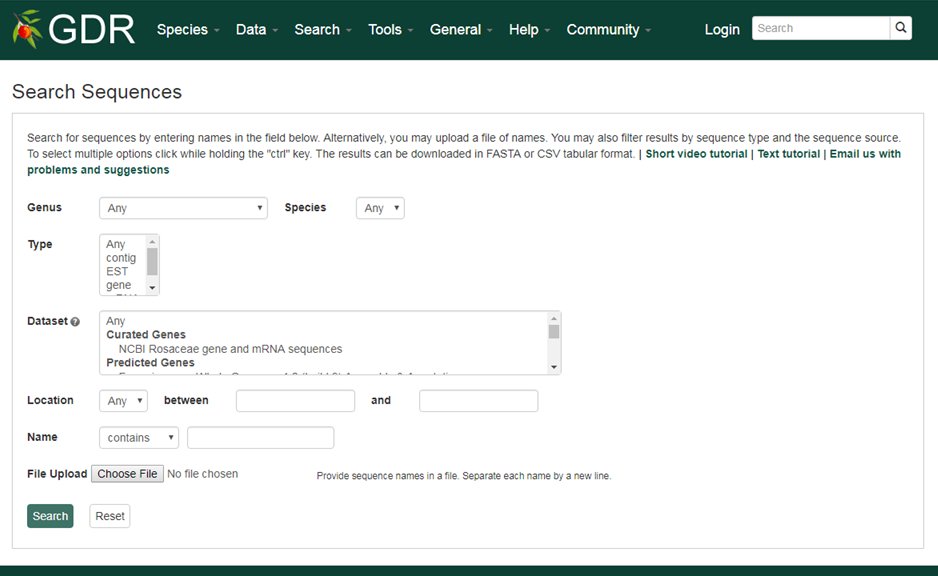
Search Sequences
If you have any questions/comments/feedback about the trait loci site, please let us know via the contact form.
Tools
To access the different Tools available on GDR, click on the 'tools' menu in the header and then select the tool you want to use. Many of the tools are also quickly accessed through links in the 'Tools Quick Start' section on the GDR homepage. To learn more about each search tool, please see the links below the figure.
How to view pan-genome data
How to view pan-genome data
To view available pangenome data click Pan-genome Data under Data dropdown.
Project page provide information on the pan-genome project as well as hyperlinks to the various pages and tools to acces the associated data. When pan-genome graph is available for the dataset, users can access the graph using the UCSC Genome Browser. Open the link for 'Pan-genome graph'.
Below is a snapshop of a UCSC Genome Browser showing pan-genome graph. Learn what the colors mean and some other useful hints.
JBrowse
GDR has an instance of the JBrowse genome browser for viewing genome data. A list of the genomes available in GDR can be accessed by clicking the JBrowse link in the Tools menu. Please watch the JBrowse tutorial for more details about how to navigate and use JBrowse.
Sequence Retrieval
The Sequence Retrieval tool allows downloading of nucleotide and protein sequences including chromosomes, scaffolds, genes, mRNAs, transcript coding sequences, protein, reftrans contigs and unigene contigs. For the sequences aligned to larger sequences, such as genes, mRNAs and transcript coding sequences, a numeric value specifying the number of upstream bases and downstream bases can be typed in the text boxes.
Below are currently available datasets for searching.
- Fragaria ananassa GDR RefTrans V1
- Fragaria Unigene v5.0
- Fragaria vesca Whole Genome v1.0 (build 8) Assembly & Annotation
- Fragaria vesca Whole Genome v1.1 Assembly & Annotation
- Fragaria vesca Whole Genome v4.0.a1 Assembly & Annotation
- Malus Unigene v5.0
- Malus x domestica GDDH13 v1.1 Whole Genome Assembly & Annotation
- Malus x domestica GDR RefTrans V1
- Malus x domestica Whole Genome v1.0 Assembly & Annotation
- Malus x domestica Whole Genome v1.0p Assembly & Annotation
- Prunus avium GDR RefTrans V1
- Prunus avium Whole Genome Assembly v1.0 & Annotation v1 (v1.0.a1)
- Prunus persica GDR RefTrans V1
- Prunus persica Whole Genome Assembly v2.0 & Annotation v2.1 (v2.0.a1)
- Prunus persica Whole Genome v1.0 Assembly & Annotation
- Prunus Unigene v5.0
- Pyrus communis Genome v1.0 Draft Assembly & Annotation
- Pyrus Unigene v5.0
- Rosaceae Family Unigene v5.0
- Rosa chinensis Whole Genome v1.0 Assembly & Annotation
- Rosa Unigene v5.0
- Rubus GDR RefTrans V1
- Rubus GDR RefTrans V2
- Rubus occidentalis Whole Genome Assembly v1.0 & Annotation v1
- Rubus occidentalis Whole Genome Assembly v1.1
- Rubus Unigene v5.0
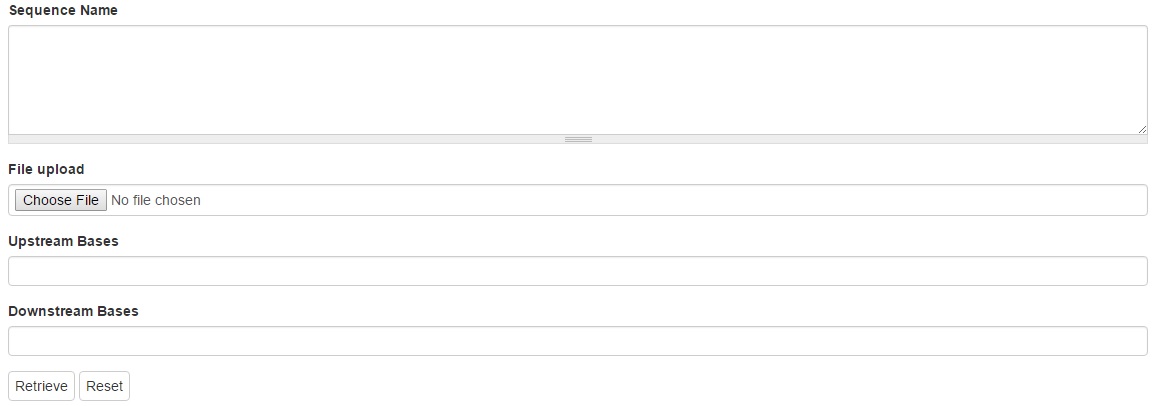
1. Sequence Name
The names of the sequences to be retrieved can be typed in the text box. Each name should be separated with a new line or comma. Leave blank to retrieve all features matching other criteria.
2. Upstream and downstream bases
A numeric value specifying the number of upstream bases and downstream bases to include in the downloaded sequences can be typed in the text boxes. This only works if the feature is aligned to a larger sequence.
If you have any questions/comments/feedback about the sequence retreival tool, please let us know via the contact form.
Synteny Viewer
Viewing RNA Expression Data
GDR has the Tripal Analysis Expression module to view differential expression values from RNA-seq experiments. Information about the different expression datasets on GDR is displayed in Expression Heatmap page. Expression data for the genome associated mRNAs or assembled transcriptome contigs is visualized either on the associated gene or mRNA page, or by creating a heatmap.
To view if your gene or mRNA of interest have associated expression data, search and find them in "Search Gene and Transcripts". Genes of mRNAs that have expression data will have an "Expression" category on the left hand menu of the feature page.
IMAGE
When "Expression" is clicked, the center pane will change to display the available expression data. Hovering over the bars on the chart displays the expression value and more infomation about the sample. Clicking on the bars will open the information page for the biomaterial, which has links to the analysis details page. There are also options to sort or edit the display of the expression data. When the gene/mRNA is associated with more than one analysis page, a dropdown will show up so that users can change the analysis.
GDR Video Tutorials
Please click on the tutorial videos for our databases on the MainLab YouTube channel. The searches and tools use the same framework across all of our databases. New videos are released every month except the months we send out quarterly newsletters. The newest video and links to key ones are featured below.
Data Searches: Trait | GWAS | Marker | Genetic Maps | QTL | Genes/Transcripts